Disease Diagnosis with Deep Learning
A symptom-checker built around a Dense Neural Network implemented from scratch in MATLAB.
Project overview. This project addresses the critical issue of medical misdiagnosis by developing an AI-powered symptom checker. To ensure a deep understanding of the underlying mathematics, the Deep Neural Network (DNN) and backpropagation algorithms were implemented entirely from scratch in MATLAB, without relying on high-level deep-learning toolkits.
Introduction
Misdiagnosis is a pervasive issue in modern healthcare. Studies indicate that approximately 11% of medical problems result in a misdiagnosis, leading to nearly 795,000 instances of permanent disability or death annually in the United States alone.
The goal of this project was to create a decision-support system that can identify complex patterns between symptoms and diseases. By acting as a “second opinion,” such a system allows medical professionals (and patients) to explore a wider range of diagnostic possibilities, particularly for infrequent diseases that might otherwise be overlooked.
Methodology
1. Data preprocessing
The model was trained on a dataset of 41 diseases and 131 unique symptoms across 4920 datapoints. The raw data required significant preprocessing:
- Mapping — diseases converted to index representations; symptoms encoded as numerical features.
-
One-hot encoding — labels (diseases) one-hot encoded. For disease index 2, the label vector becomes:
\[y = [0, 1, 0, 0, \dots, 0]\] - Train/test split — randomized to ensure generalization.

Figure 1: Snapshot of the raw dataframe (left) and the list of possible diseases (right).
2. Deep learning from scratch
To master fundamentals, I avoided standard libraries (PyTorch, TensorFlow) for the core model logic. Instead I implemented the network architecture manually in MATLAB — coding the matrix operations for the forward pass and deriving the gradients for backpropagation by hand.
The network is a Dense Neural Network with:
- Input layer — 131 nodes (one per symptom)
- Hidden layers — fully connected with learnable weights and biases
- Output layer — 41 nodes (one per disease) with Softmax activation

Figure 2: The properties of the custom Neural Network class implemented from scratch.
3. Interactive application
The trained model was deployed into a MATLAB App Designer interface that lets users select symptoms from a checklist and receive a real-time diagnosis.
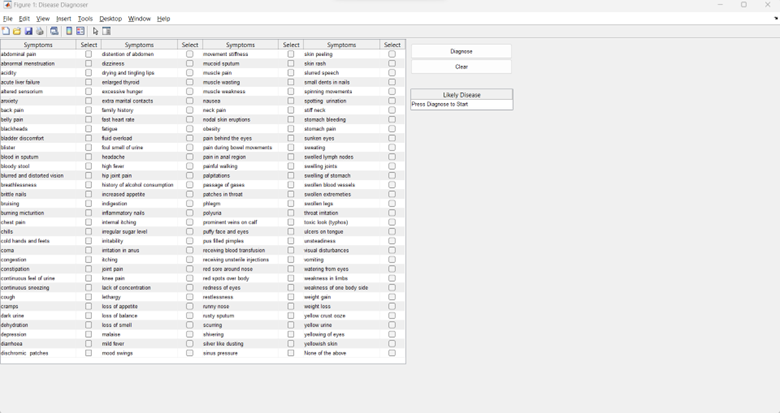
Figure 3: The UI allowing users to select symptoms.
Results and discussion
Performance
The model achieved near-perfect accuracy on the testing set within the scope of the provided dataset. However, accuracy in a controlled environment can be misleading when applied to real-world medical diagnostics.
The Softmax-vs-Sigmoid dilemma
A significant finding was the limitation of using Softmax for this task. Softmax enforces that the sum of probabilities across all classes equals 1:
\[\sigma(z)_i = \frac{e^{z_i}}{\sum_{j=1}^K e^{z_j}}\]This setup treats the problem as multi-class classification (the patient has exactly one disease out of 41) rather than multi-label classification (the patient could have comorbidities, or none of the above). To mitigate, I implemented a 70% confidence threshold: predictions below it are discarded.
Sensitivity issues
Due to thresholding and dataset structure, the model sometimes required an excessive number of symptoms to trigger a diagnosis.
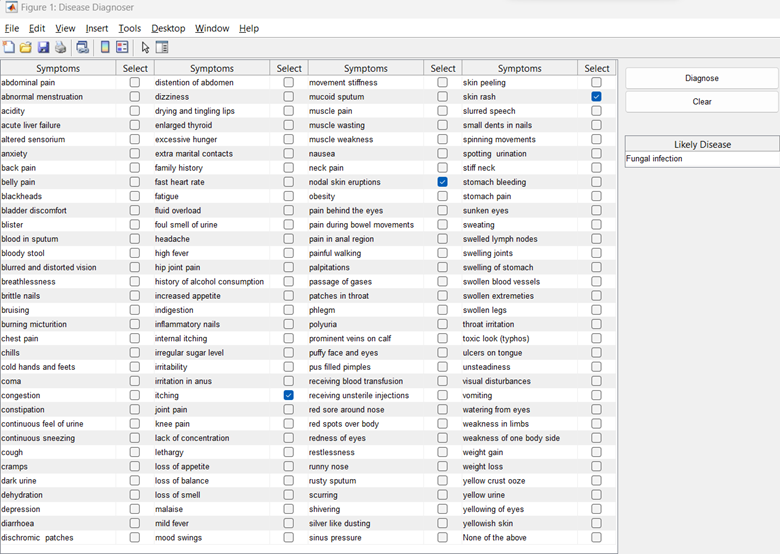
Figure 4: Diagnosing the common cold required selecting over 10 specific symptoms to cross the confidence threshold.
Key takeaways
- Fundamentals matter. Implementing backpropagation and layer logic from scratch provided deeper intuition than using high-level APIs.
- Architecture choice. For medical diagnosis, a multi-label approach (Sigmoid per node) is superior to multi-class (Softmax) — it allows independent probabilities per disease.
- Data limitations. Real-world deployment would require a vastly larger, more diverse dataset and a probabilistic framework that accounts for rare conditions.